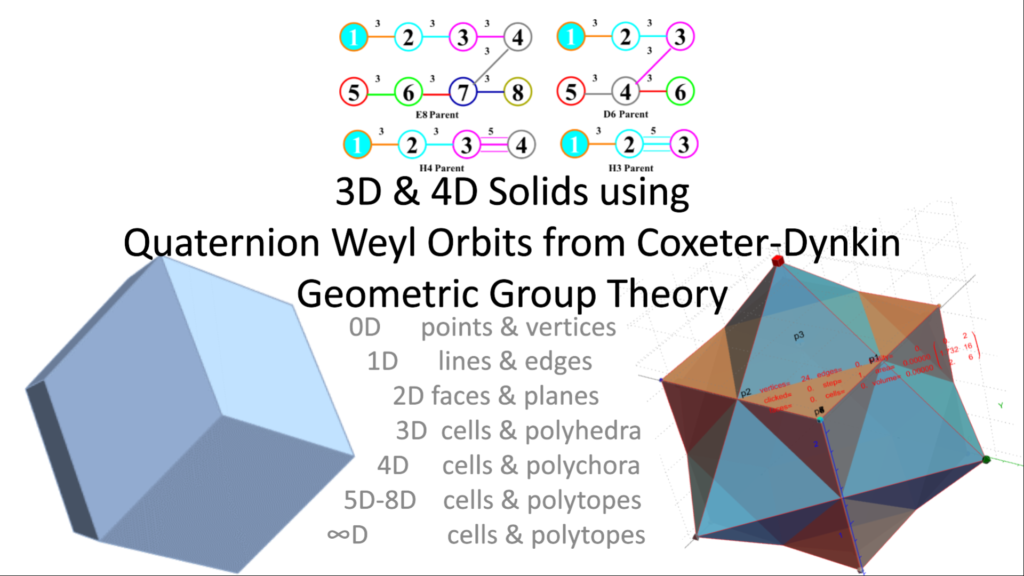
Please see this Powerpoint for an interactive presentation (160 Mb).

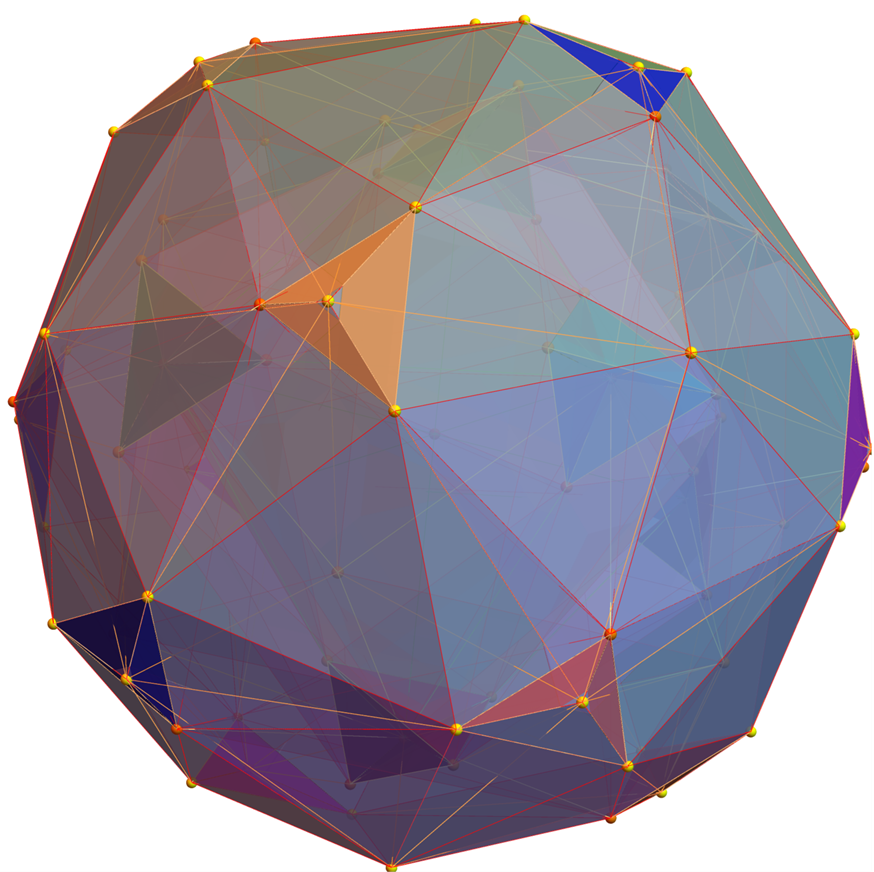
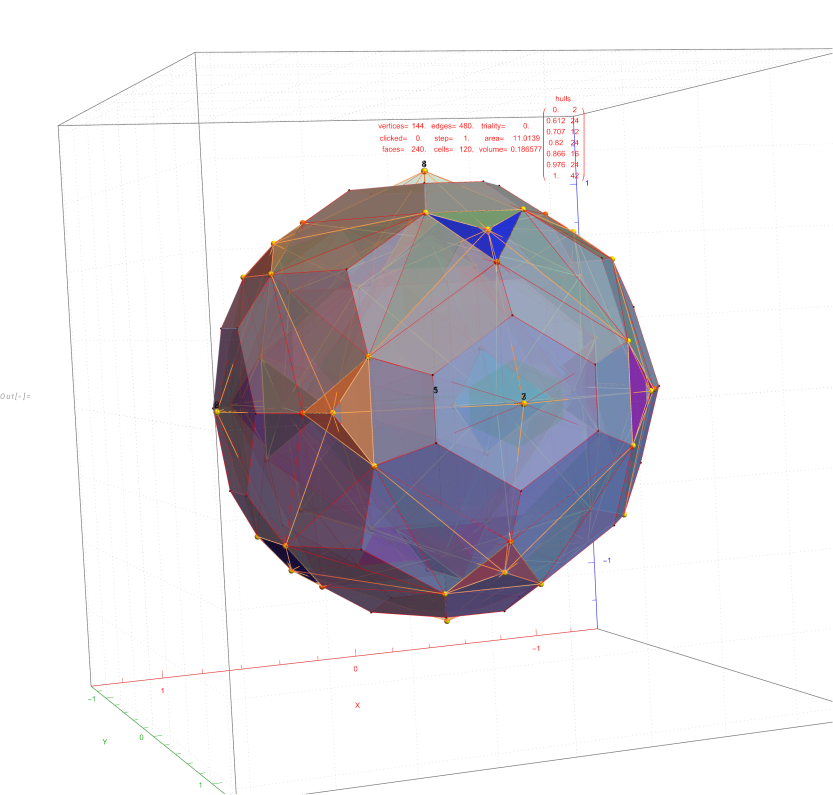
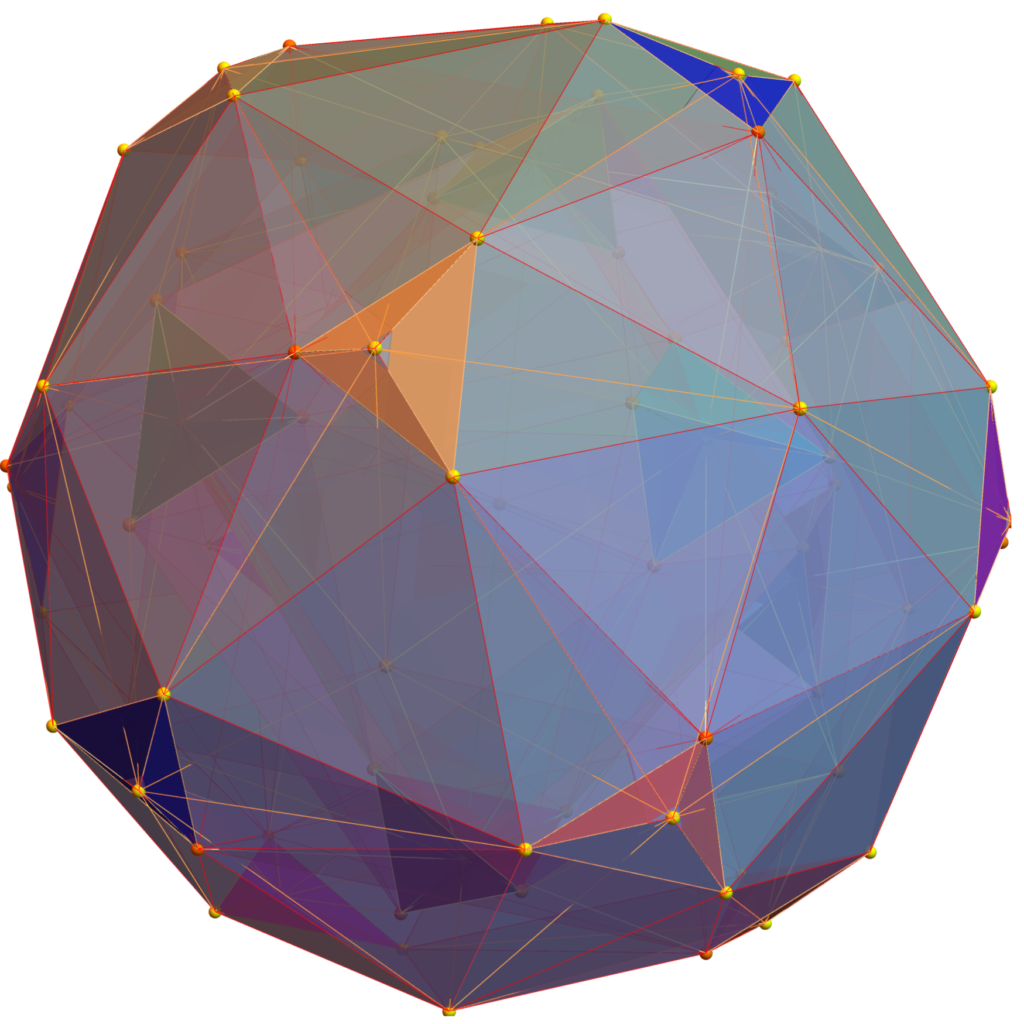
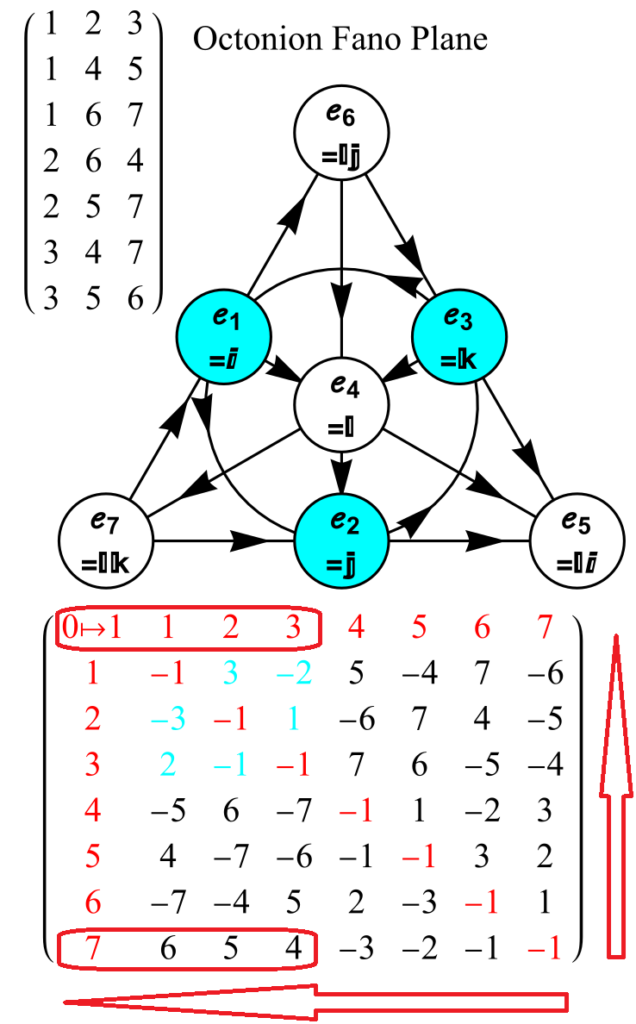
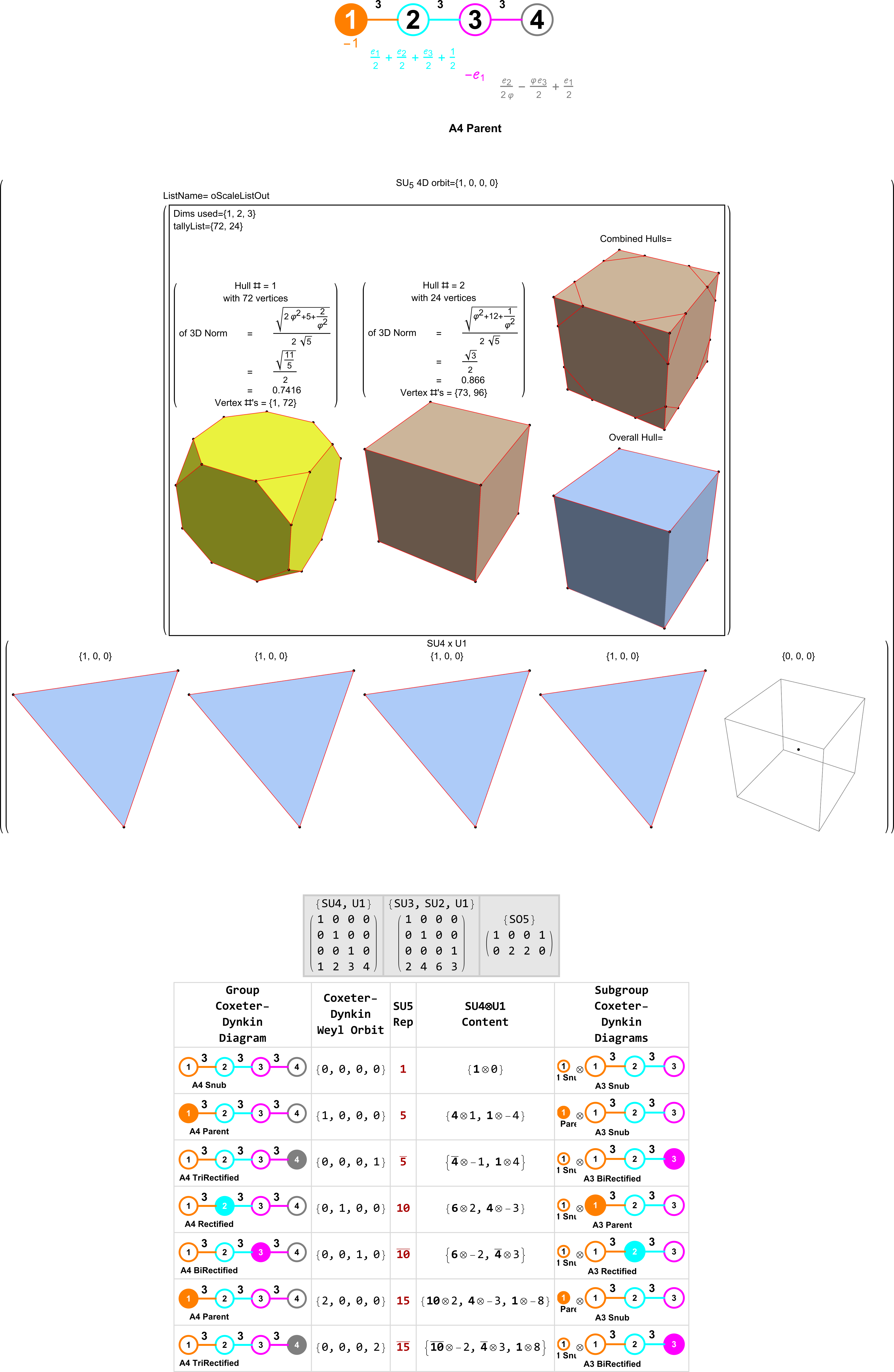
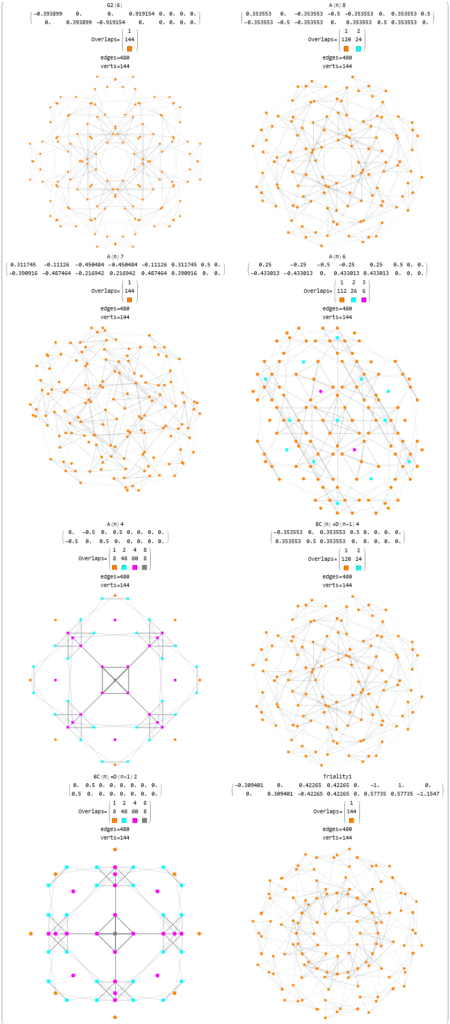
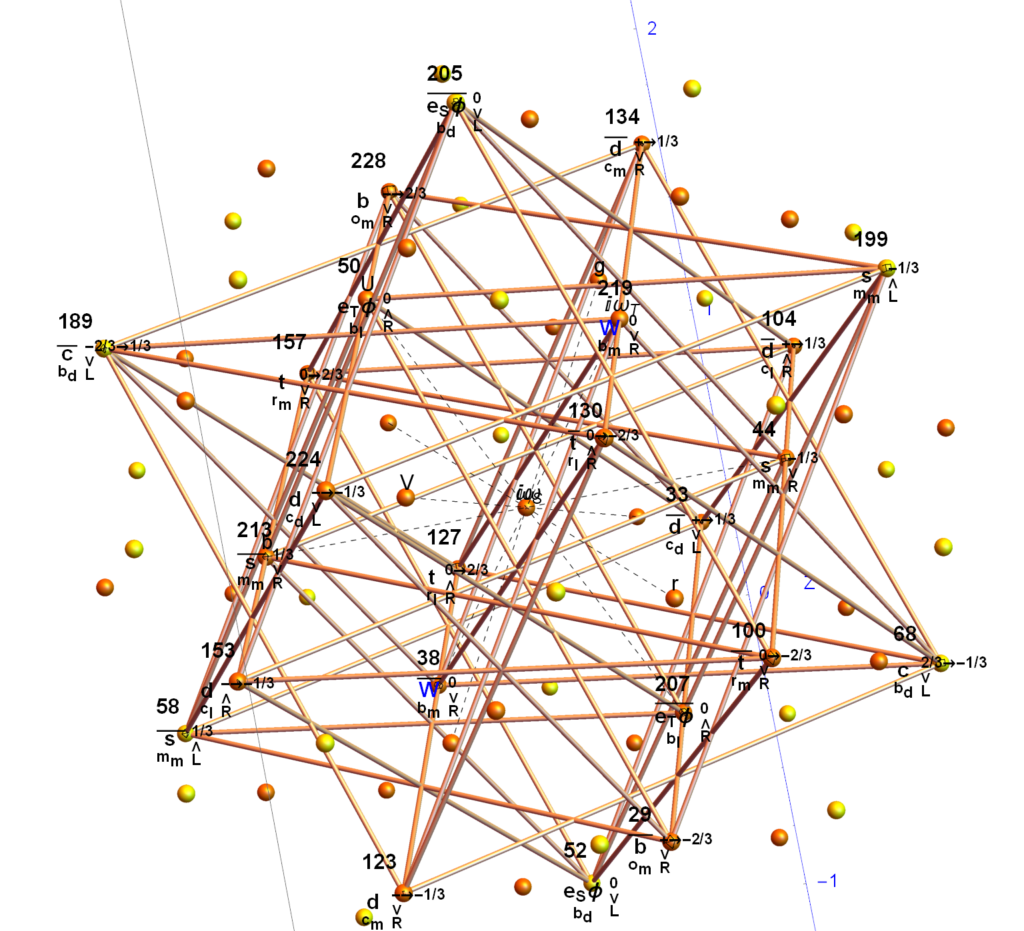
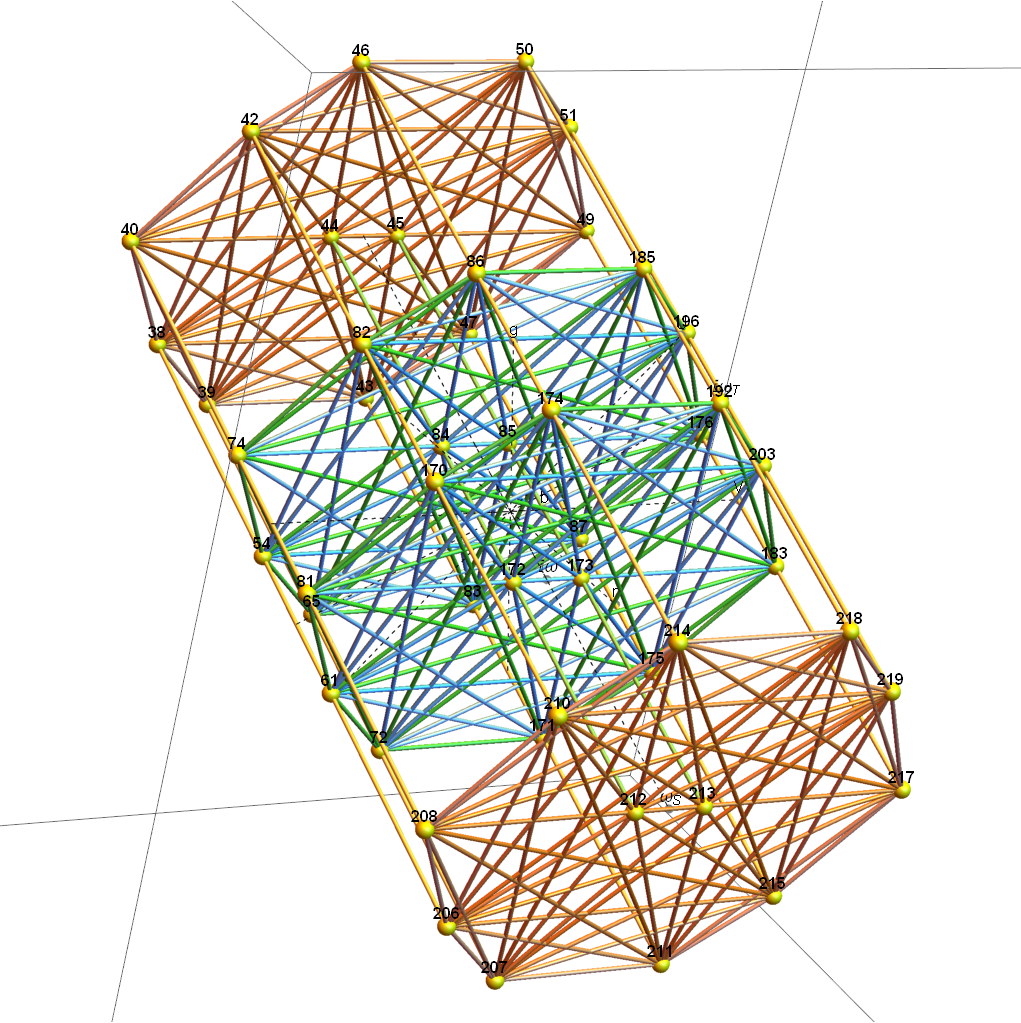
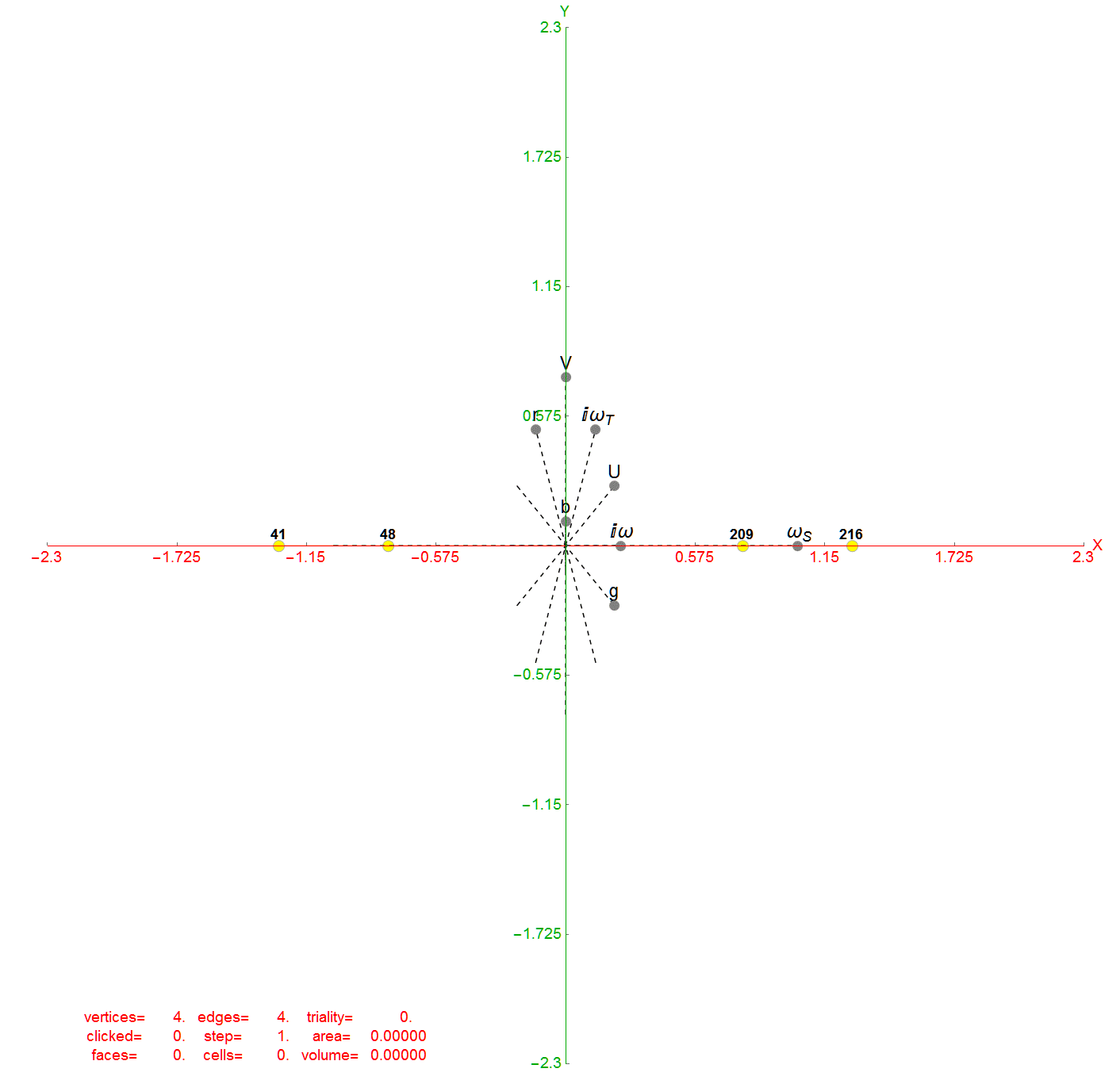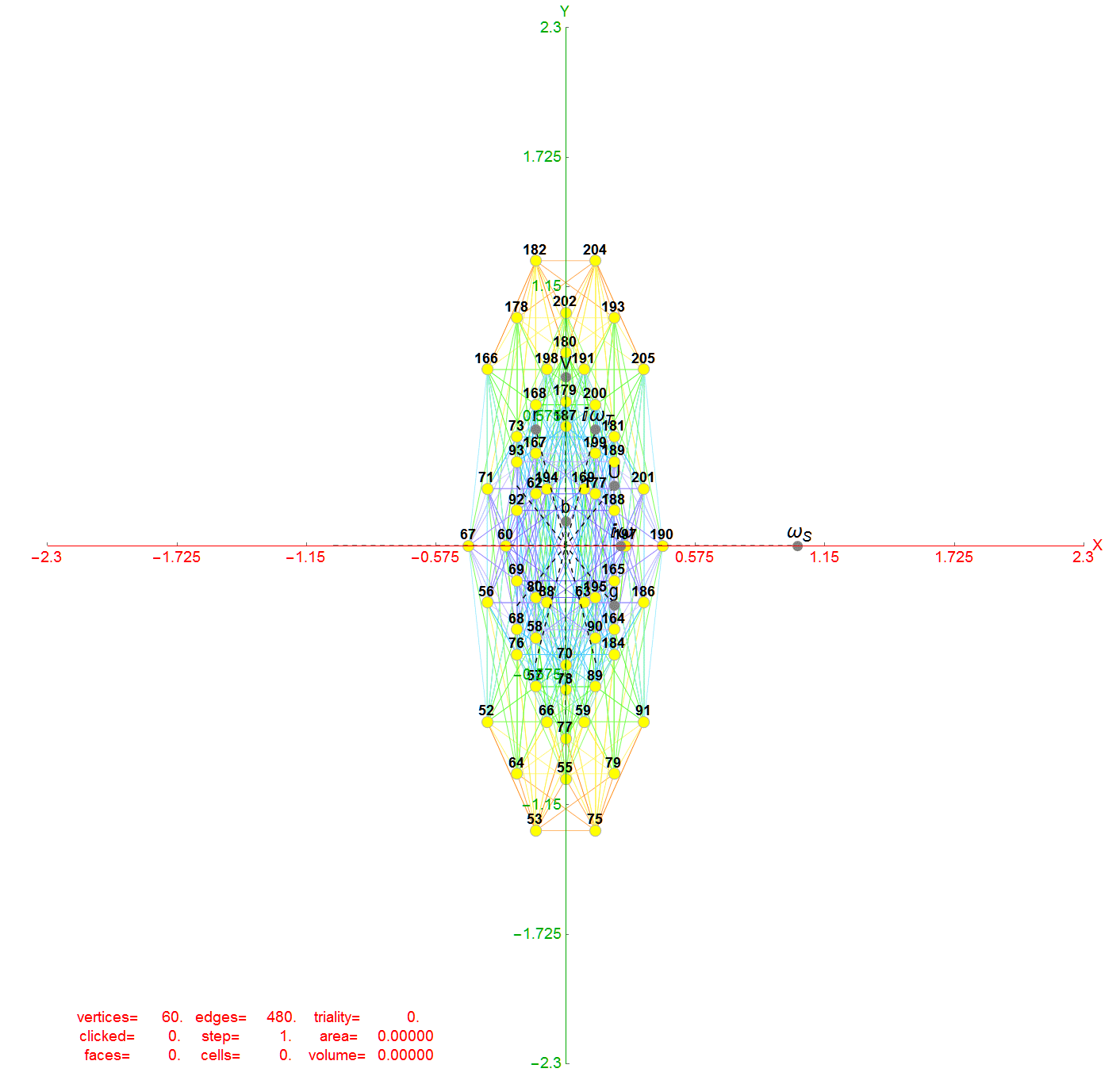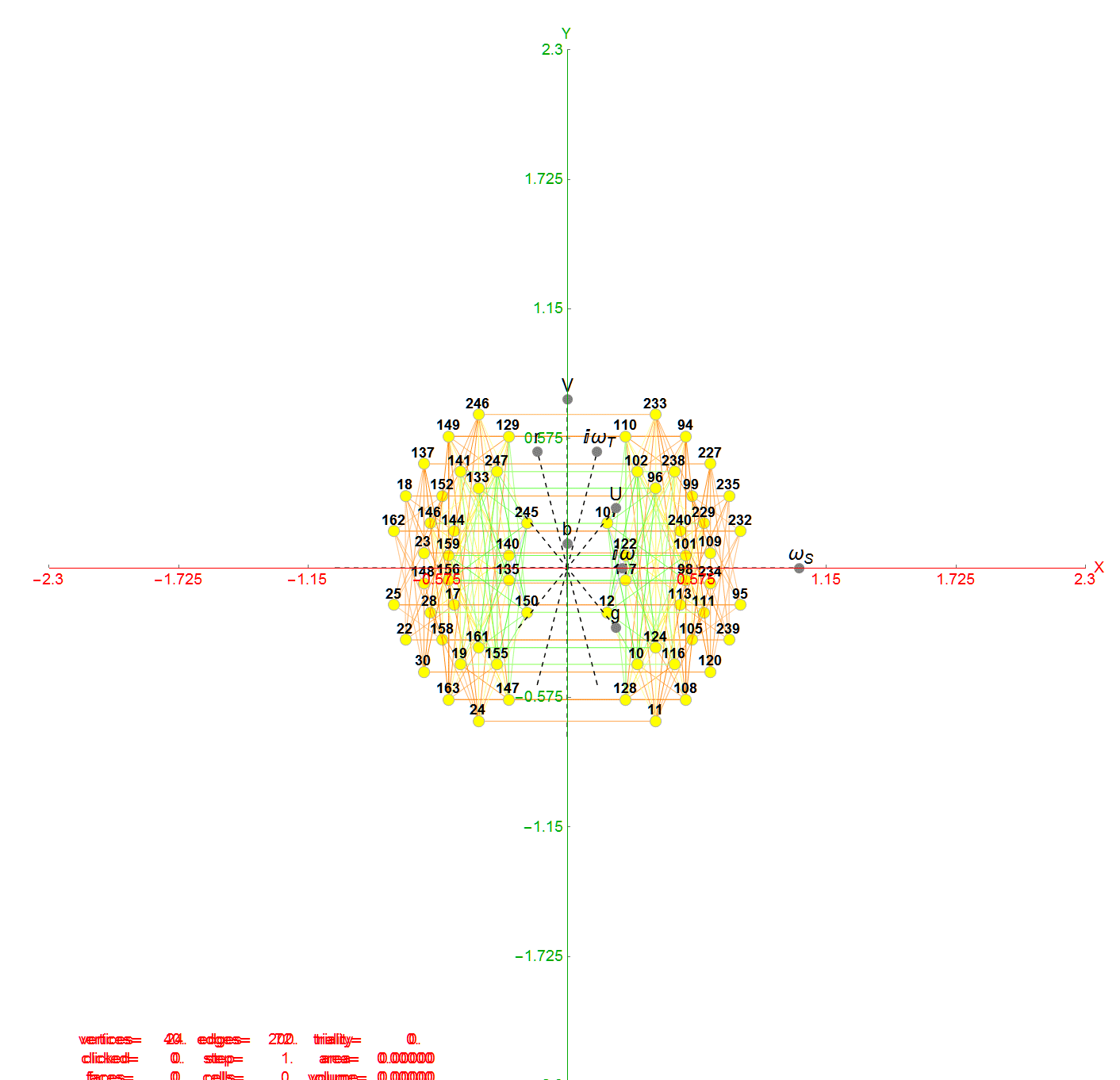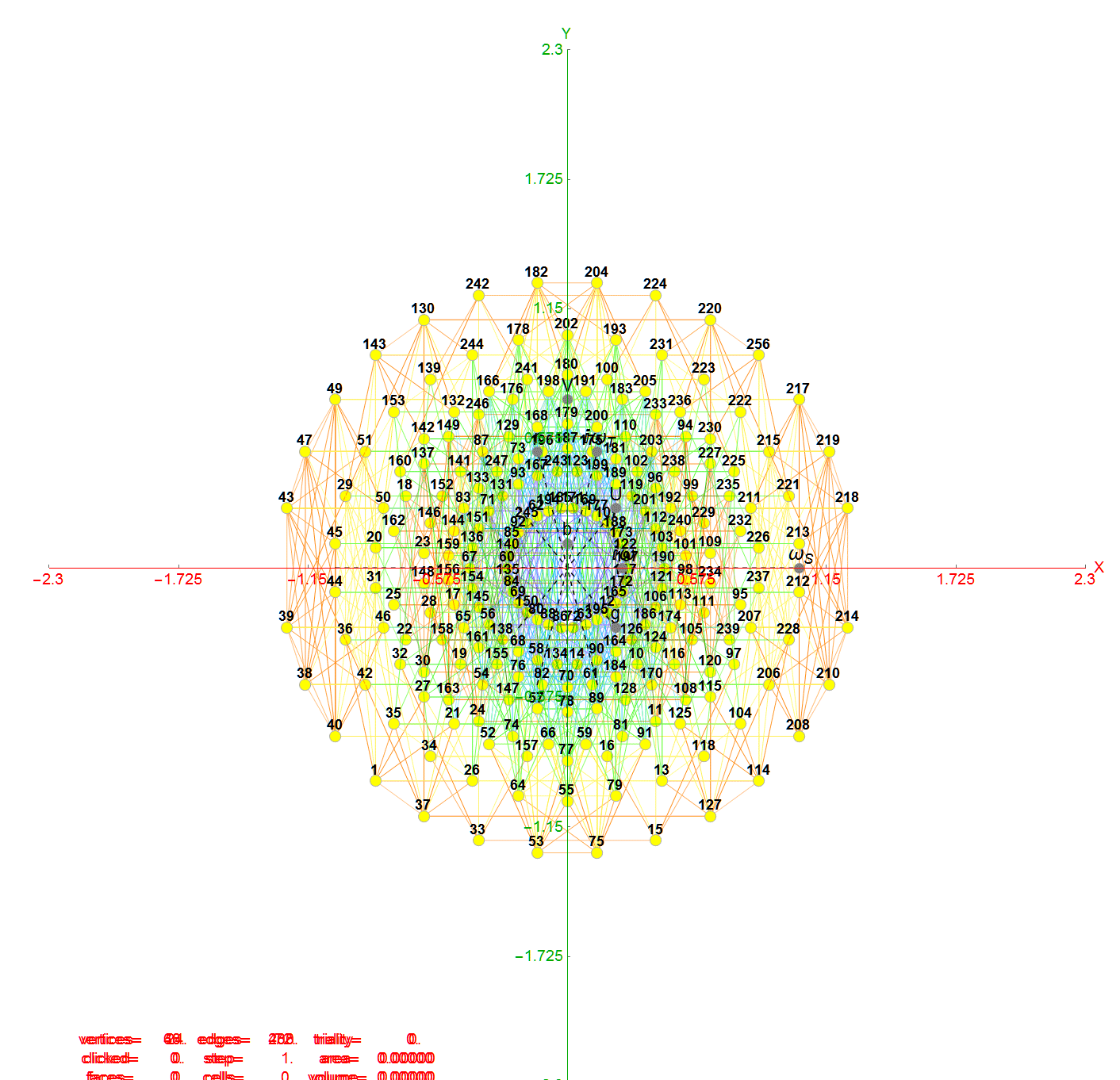
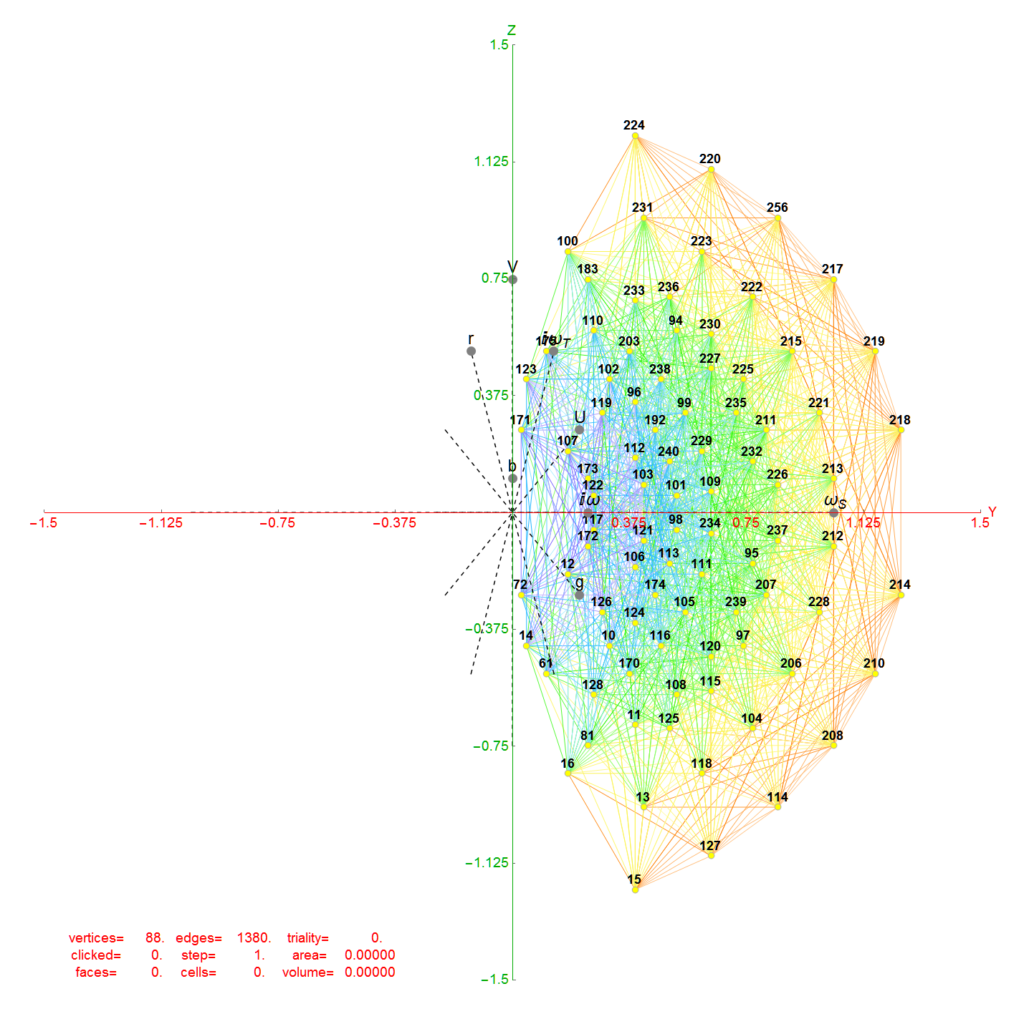
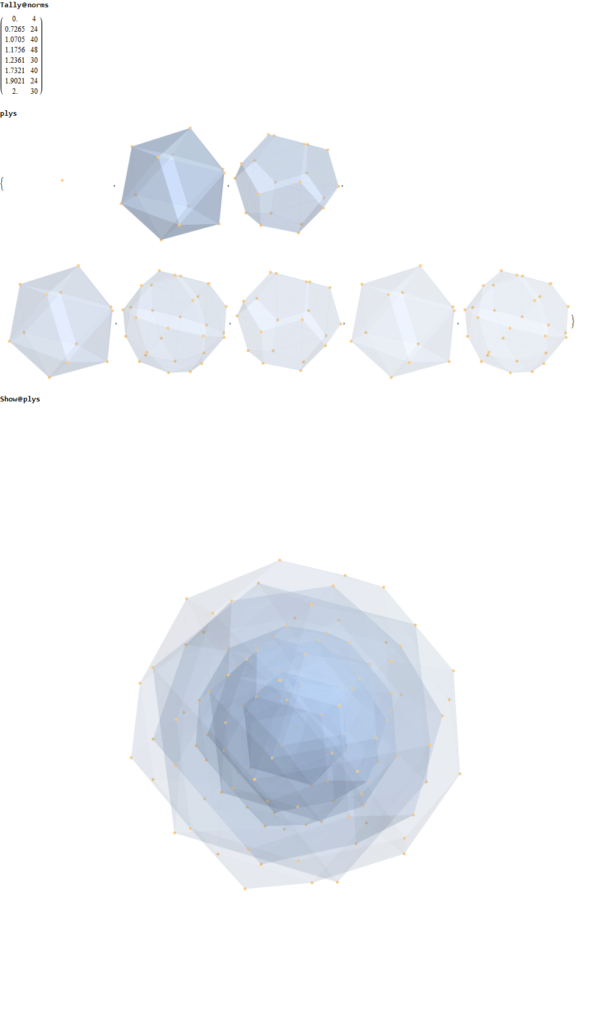
Below (and here and here in PDF and Mathematica Notebook form or here on the Cloud) are the I & I’ 600-cells based on quaternion construction from the T & T’ 24-cells. These are also documented in Coxeter’s book “Regular Polytopes” as {3,3,5} vertex first & cell first “sections” in Table V (respectively) vs. the projections shown here with one of the four dimensions projected to 0 such that the hulls are palindromic pairs of sections (e.g. for 9 sections, section 1-4 pairs with 9-6 in vertex counts equal to hull vertex counts 1-4 and the center section 5 = hull 5).

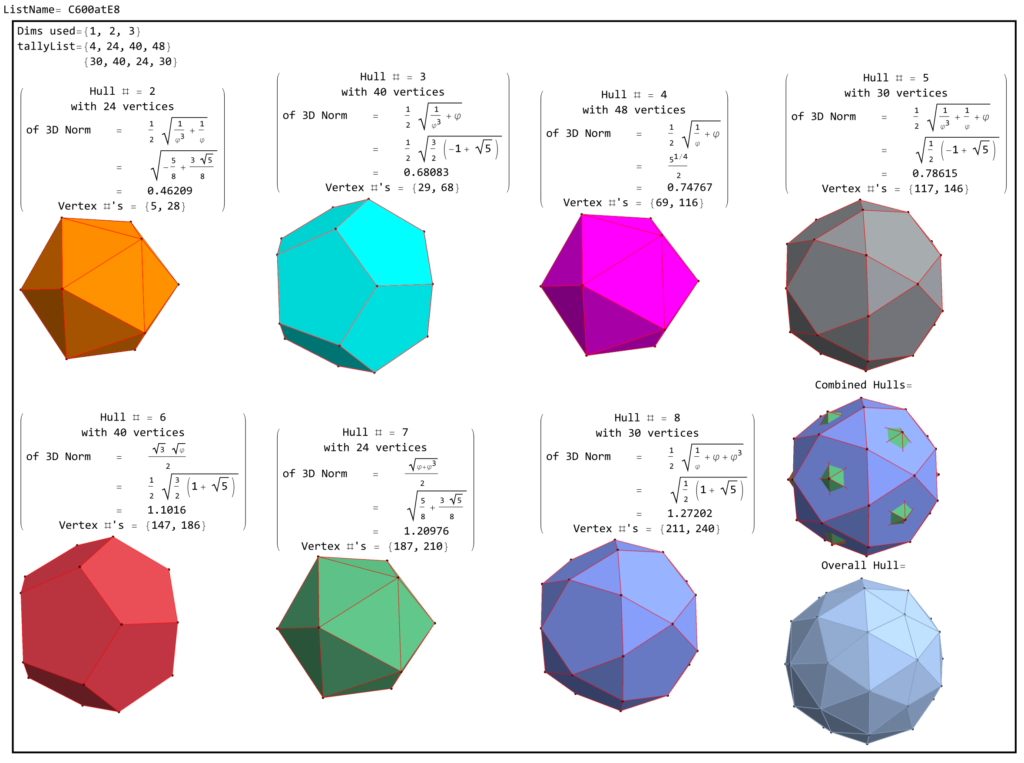
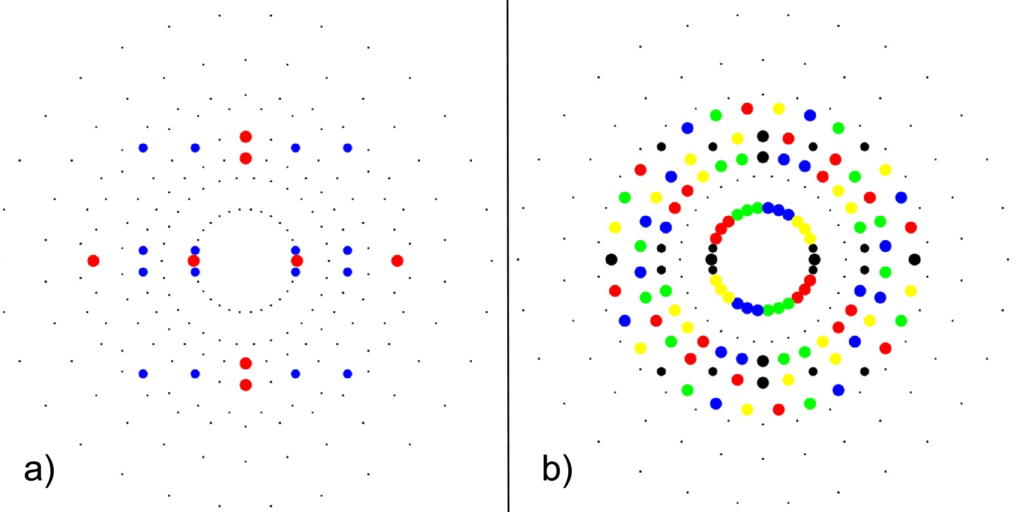
1) points at the origin
2) icosahedrons
3) dodecahedrons
4) icosahedrons
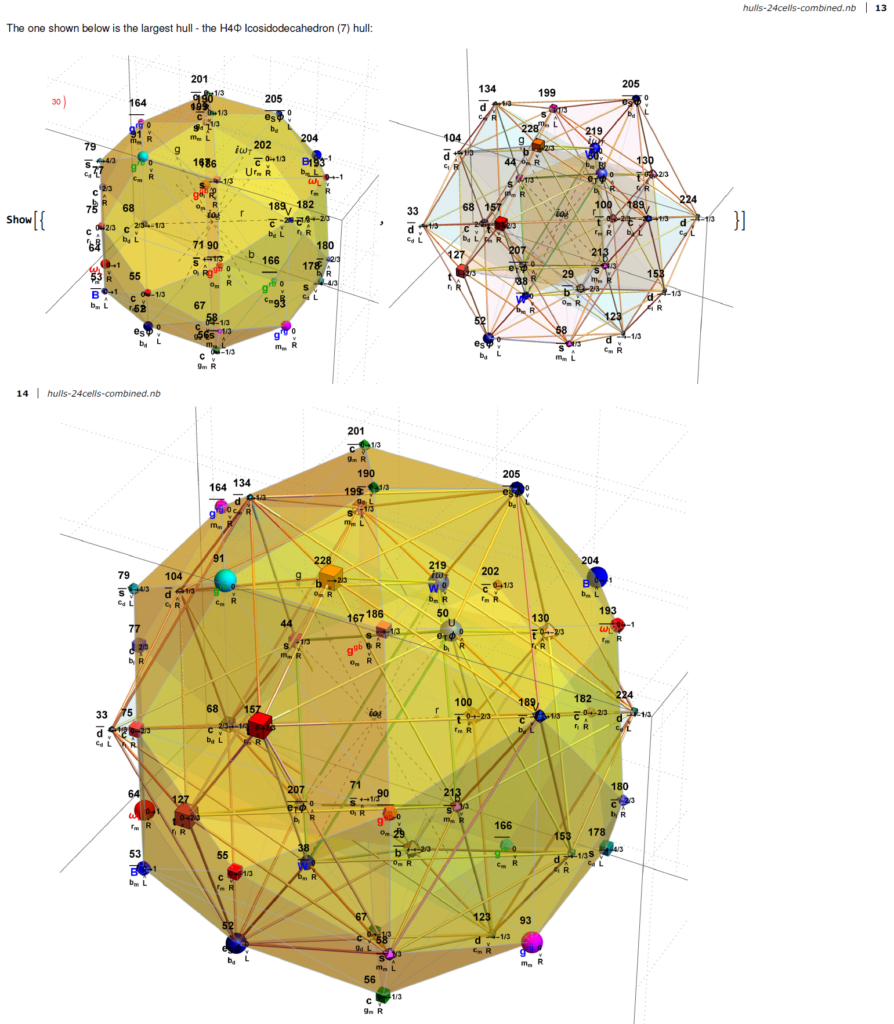
5) and a single icosadodecahedron
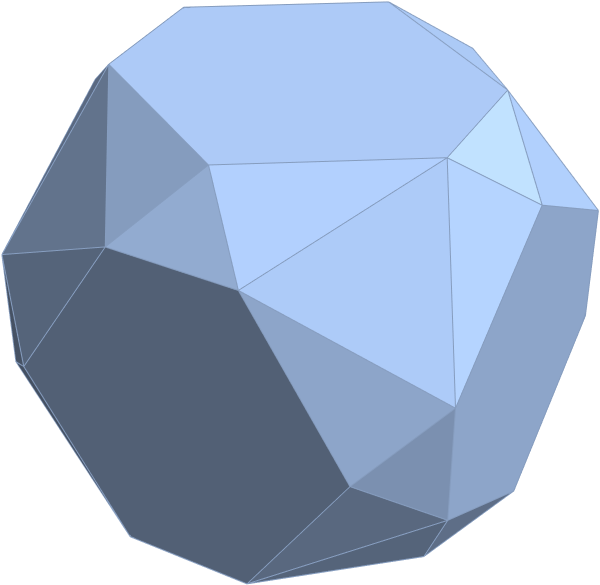
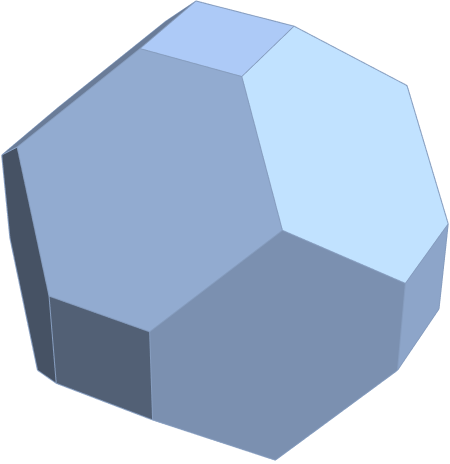
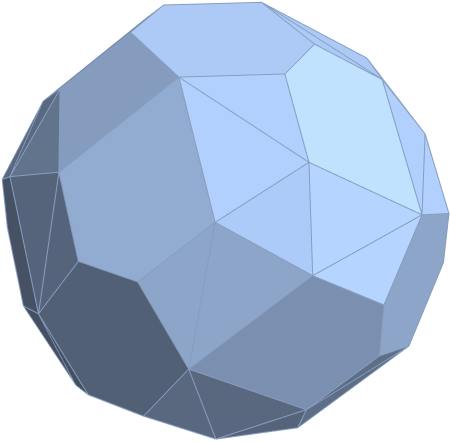
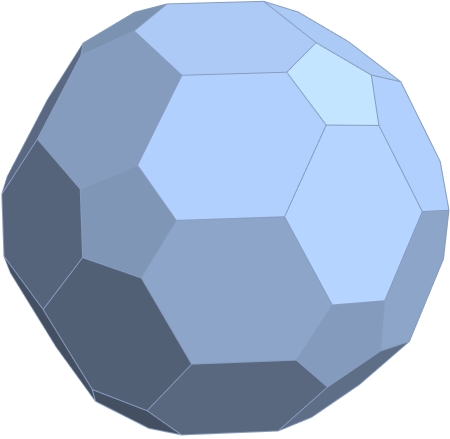
for a total of 120 vertices and an overall outer 3D hull of the Pentakis Icosidodecahedron.

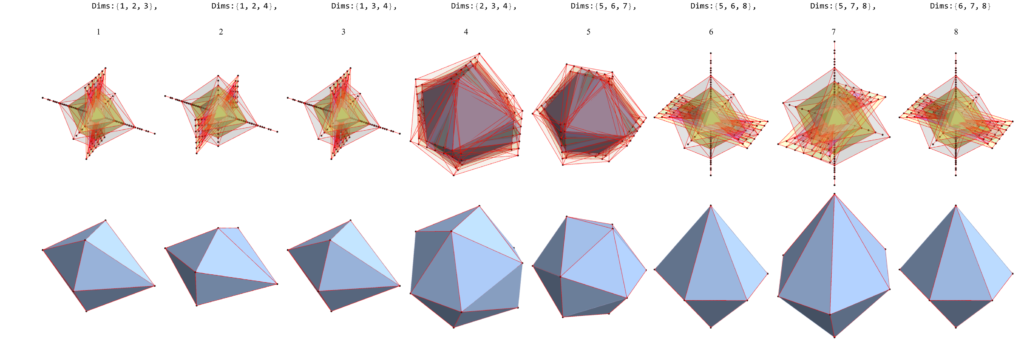
1) a pair of tetrahedrons (or single cube)
2) a pair of tetrahedrons (or single cube)
3) a pair of icosahedrons
4) truncated cube
5) rhombicuboctahedron
6) rhombicuboctahedron
7) a pair of tetrahedrons (or single cube)
8) cuboctahedron
for a total of 120 vertices and an overall outer 3D hull of a near-miss of the Pentakis Icosidodecahedron.
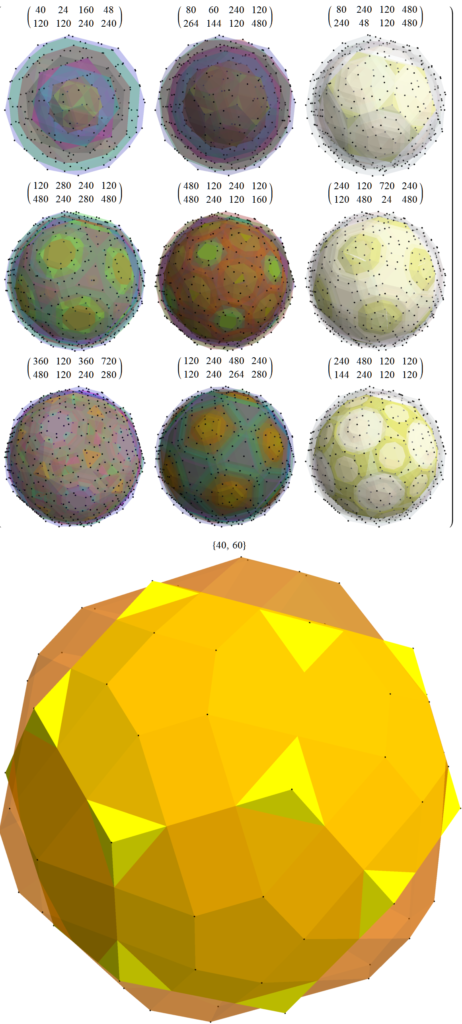
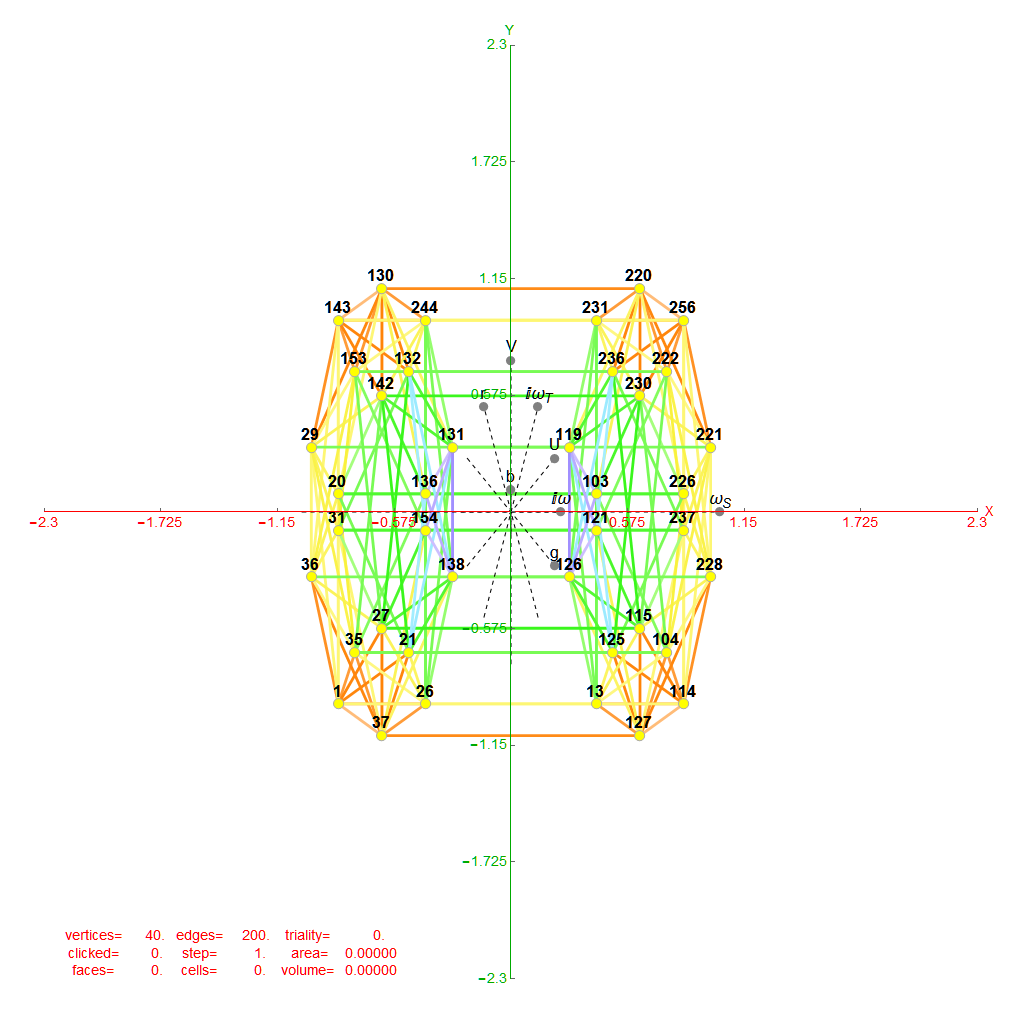
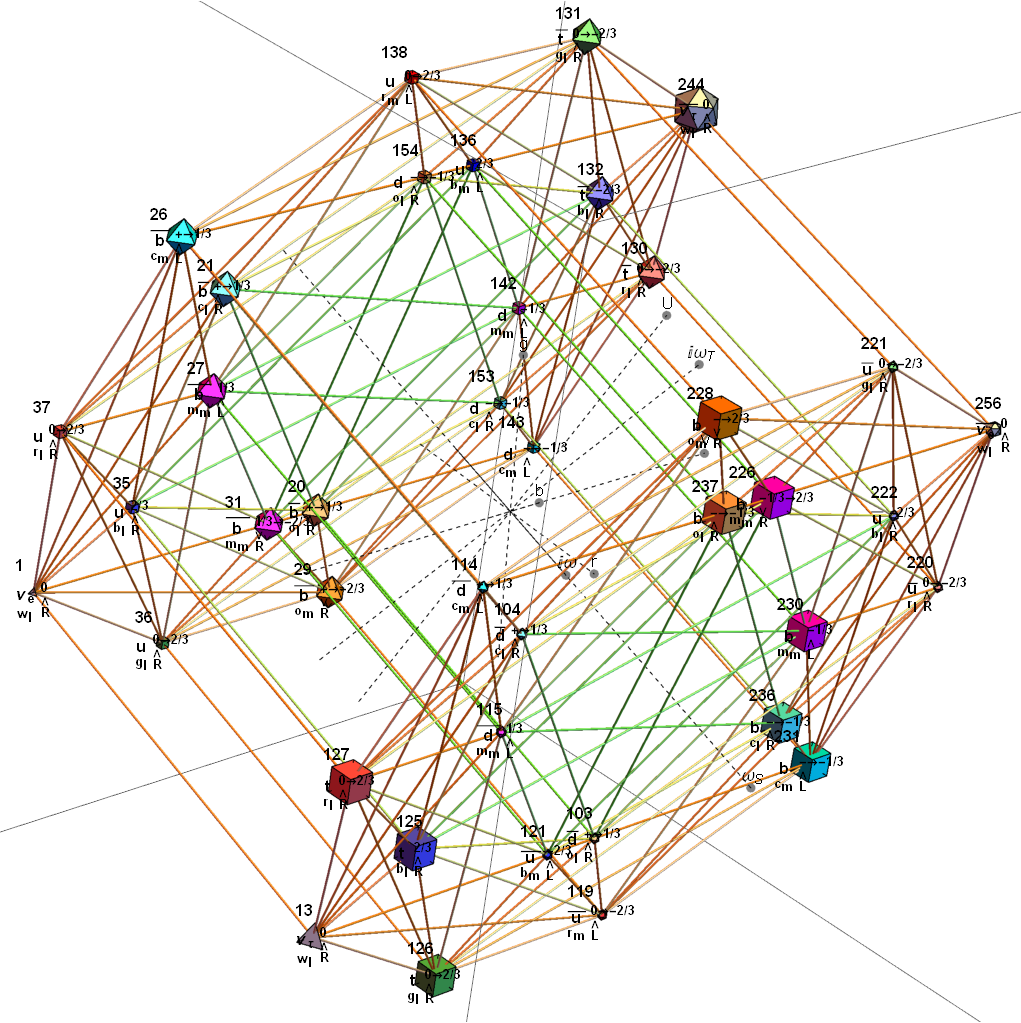
For completeness, here are the J’ (alternate) & J form of the 120-cells based on quaternion construction from the T & T’ 24-cells. These are also documented in Coxeter’s book “Regular Polytopes” as {5,3,3} vertex first & cell first “sections” in Table V (respectively) vs. the projections shown here with one of the four dimensions projected to 0 such that the hulls are palindromic pairs of sections (e.g. for 31 sections, section 0-14 pairs with 30-16 in vertex counts equal to hull vertex counts 1-15 and the center section 15 = hull 16).

Hulls 2 & 6 are cubes.
Hulls 3 & 5 are rhombicuboctahedrons.
Hulls 4 & 12 are each pairs of truncated octahedrons.
Hulls 7 & 15 are truncated cuboctahedrons.
Hull 11 is a truncated cube.
Hulls 8, 9, 10, 13, 14, and 16 are unnamed solids.

Hull 3 is a pair of icosidodecahedrons.
Hulls 4 & 5 are each pairs of truncated icosahedrons.
Hulls 6 & 8 are each rhombicosidodecahedrons.
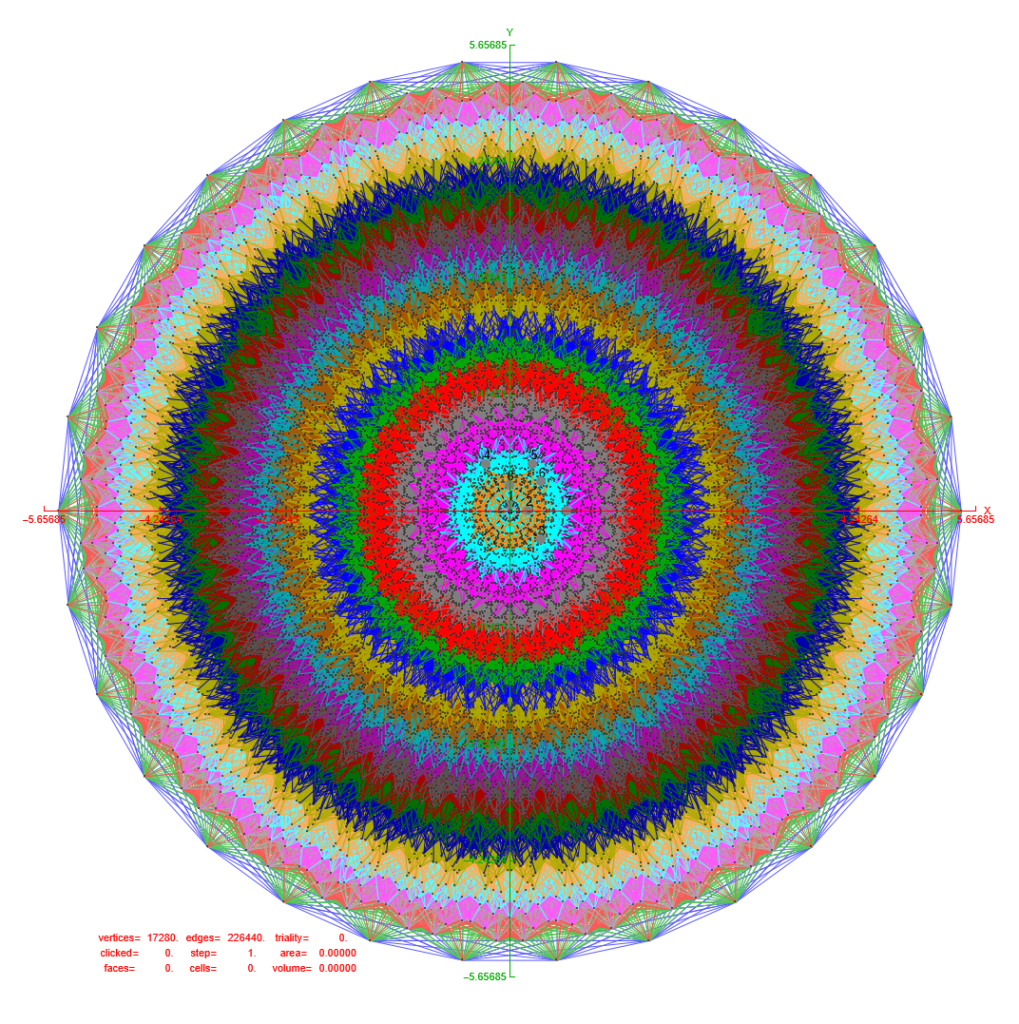
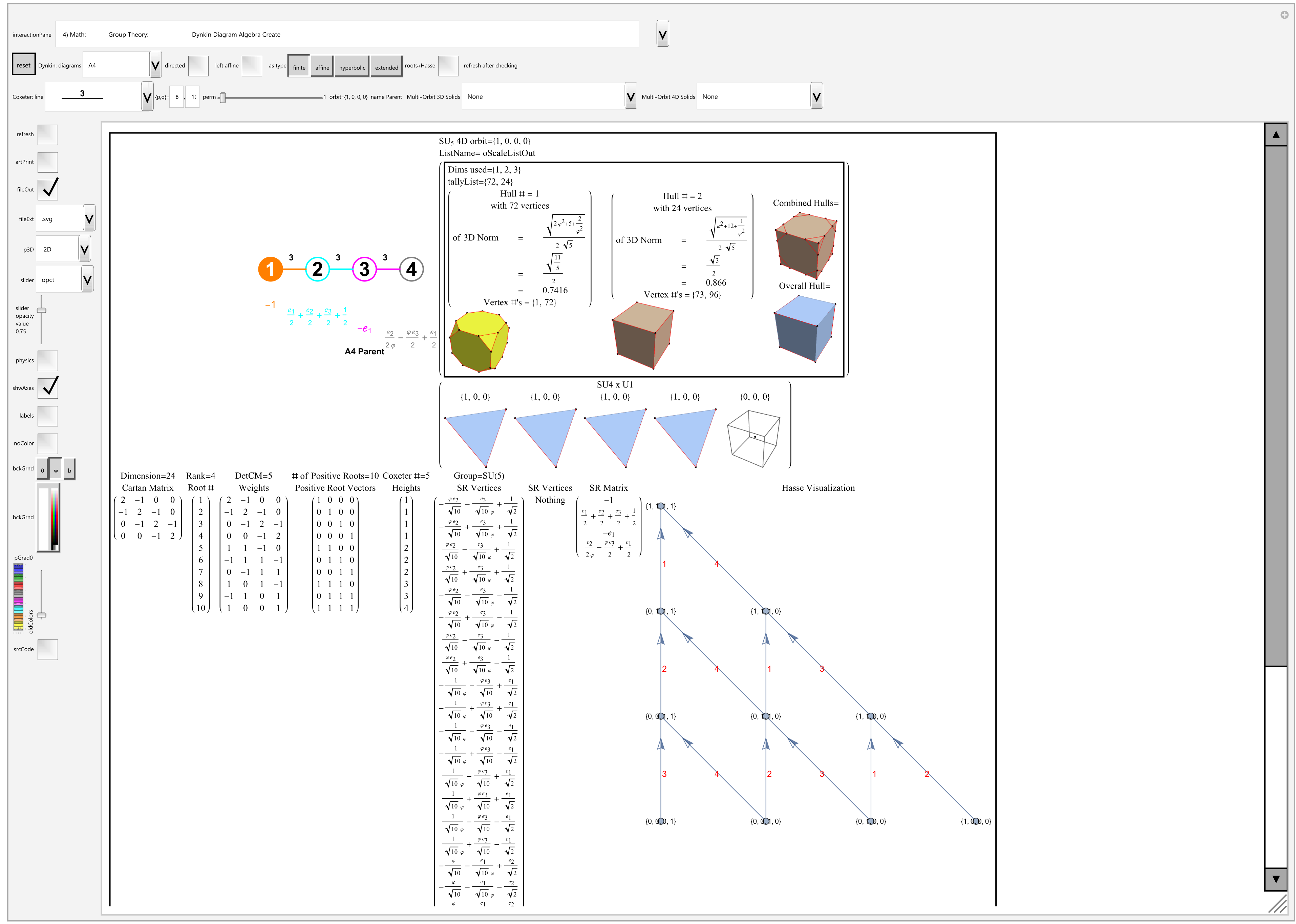
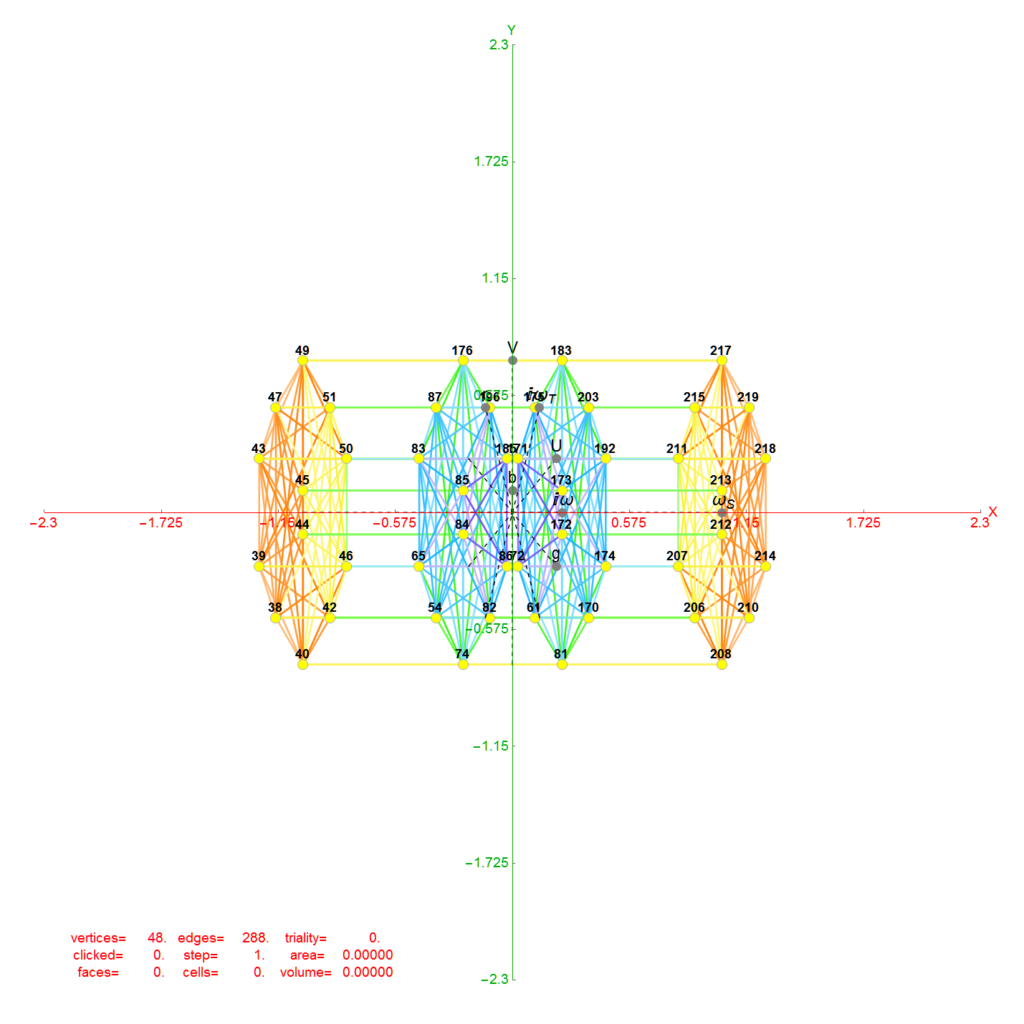
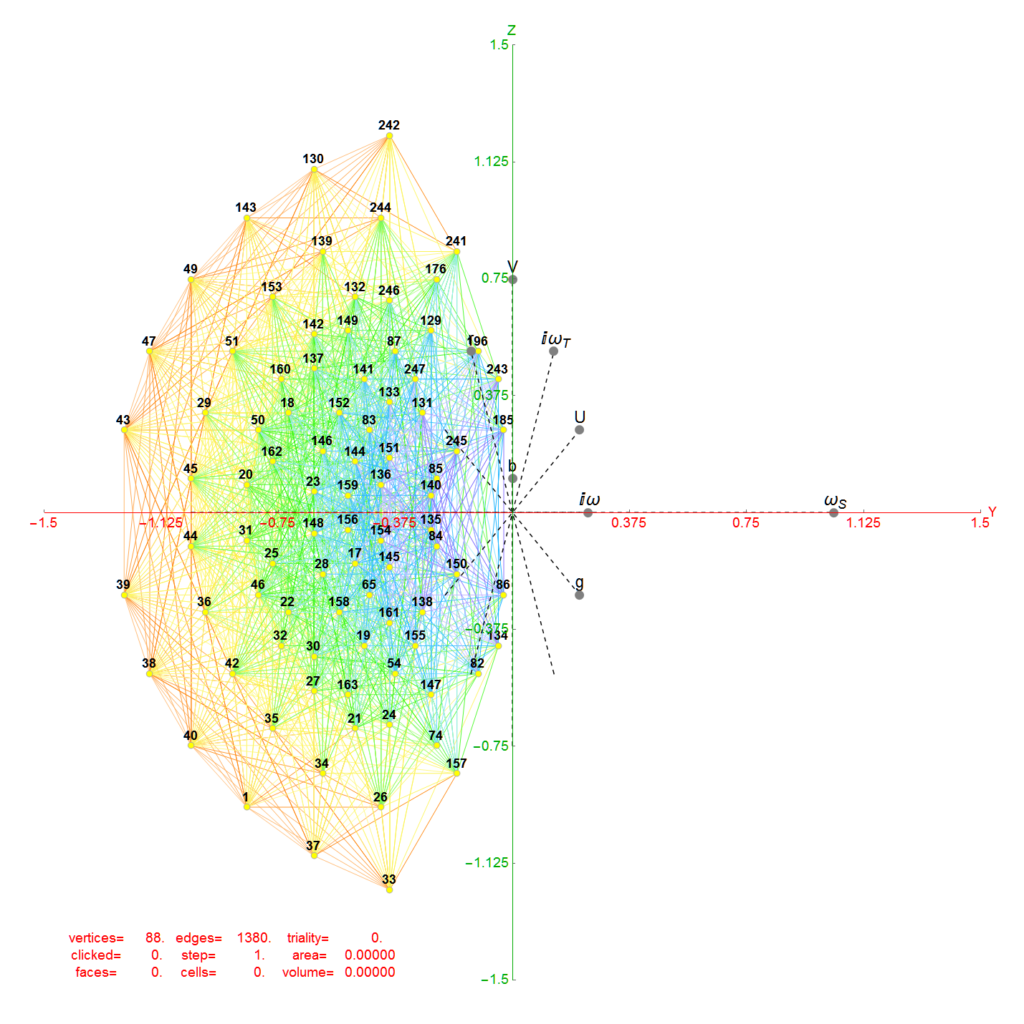
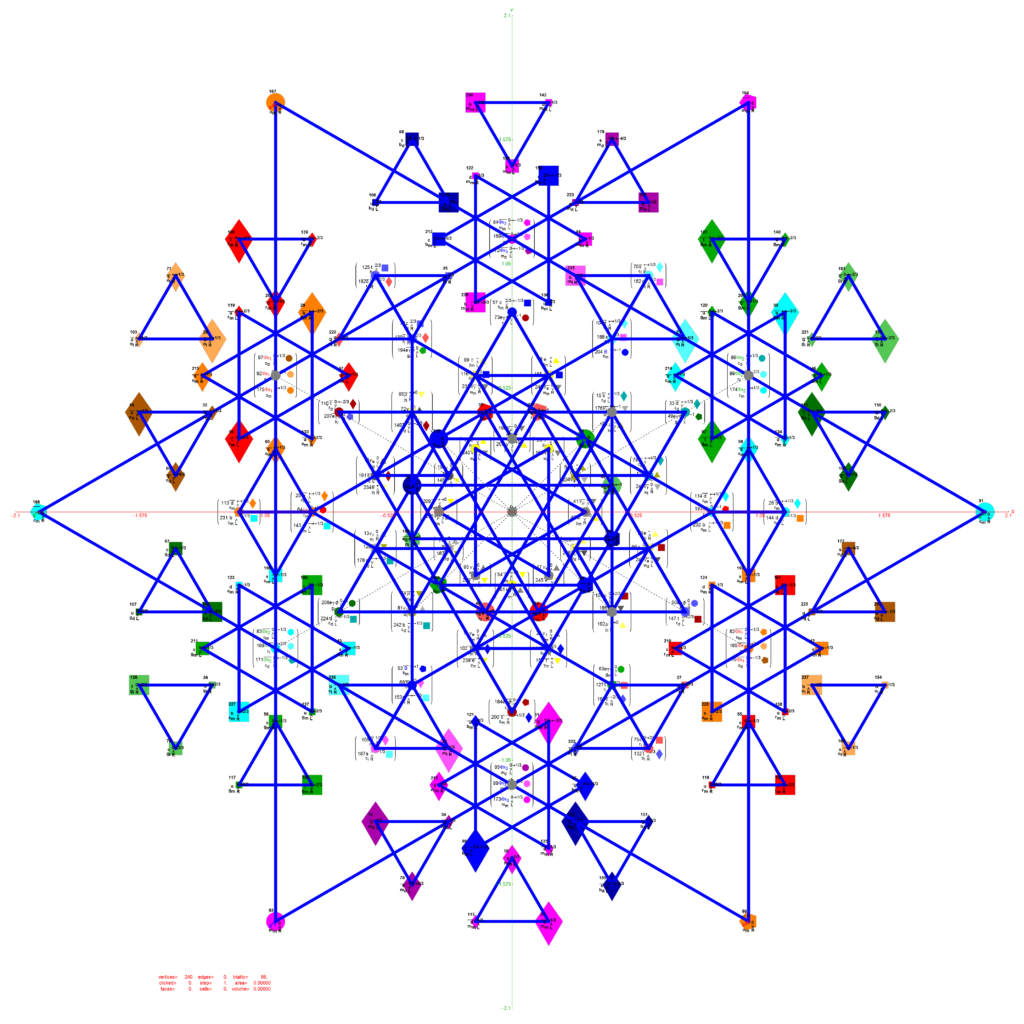
Below is an interactive Mathematica 4D visualization of the 8,16, 24, 120, and 600 cells (including alternate forms 24p, 120p, 600p). Click here or on the image below to open in a new tab.